Search
Results
Combining Biological Databases and Text Mining to support New Bioinformatics Applications

Abstract
A large amount of biological knowledge today is only available from full-text research papers. Since neither manual database curators nor users can keep up with the rapidly expanding volume of scientific literature, natural language processing approaches are becoming increasingly important for bioinformatic projects.
In this paper, we go beyond simply extracting information from full-text articles by describing an architecture that supports targeted access to information from biological databases using the results derived from text mining of research papers, thereby integrating information from both sources within a biological application.
The described architecture is currently being used to extract information about protein mutations from full-text research papers. Text mining results drive the retrieval of sequence information from protein databases and the employment of algorithmic sequence analysis tools, which facilitate further data access from protein structure databases. Complex mapping of NLP derived text annotations to protein structures allows the rendering, with 3D structure visualization, of information not available in databases of mutation annotations.
Enriching Protein Structure Visualizations with Mutation Annotations Obtained by Text Mining Protein Engineering Literature
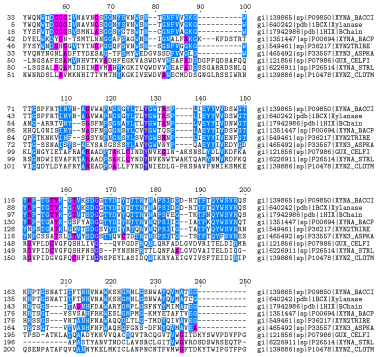

Abstract
Protein structure visualization tools render images that allow the user to explore structural features of a protein. Context specific information relating to a particular protein or protein family is not easily integrated and must be uploaded from databases or provided through manual curation of input files. We describe a mixed natural language processing and sequence analysis based approach for the retrieval of mutation specific annotations from full text articles for rendering with protein structures.
Keywords
Text Mining, Protein Structure Annotation, Protein Function, ProSAT, Xylanase
Mutation mining - A prospector's tale
Abstract
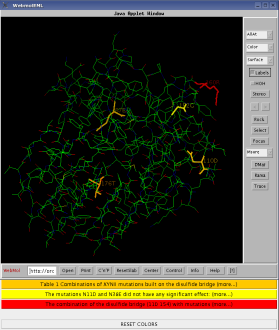
Protein structure visualization tools render images that allow the user to explore structural features of a protein. Context specific information relating to a particular protein or protein family is, however, not easily integrated and must be uploaded from databases or provided through manual curation of input files. Protein Engineers spend considerable time iteratively reviewing both literature and protein structure visualizations manually annotated with mutated residues. Meanwhile, text mining tools are increasingly used to extract specific units of raw text from scientific literature and have demonstrated the potential to support the activities of Protein Engineers.
The transfer of mutation specific raw-text annotations to protein structures requires integrated data processing pipelines that can co-ordinate information retrieval, information extraction, protein sequence retrieval, sequence alignment and mutant residue mapping. We describe the Mutation Miner pipeline designed for this purpose and present case study evaluations of the key steps in the process. Starting with literature about mutations made to protein families; haloalkane dehalogenase, bi-phenyl dioxygenase, and xylanase we enumerate relevant documents available for text mining analysis, the available electronic formats, and the number of mutations made to a given protein family. We review the efficiency of NLP driven protein sequence retrieval from databases and report on the effectiveness of Mutation Miner in mapping annotations to protein structure visualizations. We highlight the feasibility and practicability of the approach.
Keywords
Text mining - Protein structure annotation - Protein mutation - Data mining - Haloalkane dehalogenase - Biphenyl dioxygenase - Xylanase
Ontology-Based Extraction and Summarization of Protein Mutation Impact Information
Introduction
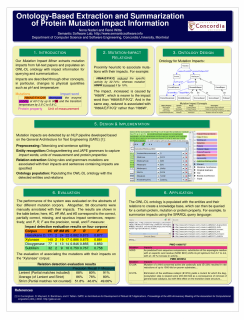 Poster at BioNLP 2010: Ontology-Based Extraction and Summarization of Protein Mutation Impact InformationNLP methods for extracting mutation information from the bibliome have become an important new research area within bio-NLP, as manually curated databases, like the Protein Mutant Database (PMD) (Kawabata et al., 1999), cannot keep up with the rapid pace of mutation research. However, while significant progress has been made with respect to mutation detection, the automated extraction of the impacts of these mutations has so far not been targeted. In this paper, we describe the first work to automatically summarize impact information from protein mutations. Our approach is based on populating an OWL-DL ontology with impact information, which can then be queried to provide structured information, including a summary.
Poster at BioNLP 2010: Ontology-Based Extraction and Summarization of Protein Mutation Impact InformationNLP methods for extracting mutation information from the bibliome have become an important new research area within bio-NLP, as manually curated databases, like the Protein Mutant Database (PMD) (Kawabata et al., 1999), cannot keep up with the rapid pace of mutation research. However, while significant progress has been made with respect to mutation detection, the automated extraction of the impacts of these mutations has so far not been targeted. In this paper, we describe the first work to automatically summarize impact information from protein mutations. Our approach is based on populating an OWL-DL ontology with impact information, which can then be queried to provide structured information, including a summary.
Ontology Design for Biomedical Text Mining

Abstract
Text Mining in biology and biomedicine requires a large amount of domain-specific knowledge. Publicly accessible resources hold much of the information needed, yet their practical integration into natural language processing (NLP) systems is fraught with manifold hurdles, especially the problem of semantic disconnectedness throughout the various resources and components. Ontologies can provide the necessary framework for a consistent semantic integration, while additionally delivering formal reasoning capabilities to NLP.
In this chapter, we address four important aspects relating to the integration of ontology and NLP: (i) An analysis of the different integration alternatives and their respective vantages; (ii) The design requirements for an ontology supporting NLP tasks; (iii) Creation and initialization of an ontology using publicly available tools and databases; and (iv) The connection of common NLP tasks with an ontology, including technical aspects of ontology deployment in a text mining framework. A concrete application example—text mining of enzyme mutations—is provided to motivate and illustrate these points.
Keywords: Text Mining, NLP, Ontology Design, Ontology Population, Ontological NLP
Mutation Miner - Textual Annotation of Protein Structures
Abstract
Protein structure visualization tools render images that allow the user to explore structural features of a protein. Context specific information relating to a particular protein or protein family is not easily integrated and must be uploaded from databases or provided through manual curation of input files. We describe a mixed natural language processing and protein sequence analysis approach for the retrieval of mutation specific annotations from full text articles for rendering with protein structures.
Integrating Wiki Systems, Natural Language Processing, and Semantic Technologies for Cultural Heritage Data Management

Abstract
Modern documents can easily be structured and augmented to have the characteristics of a semantic knowledge base. Many older documents may also hold a trove of knowledge that would deserve to be organized as such a knowledge base. In this chapter, we show that modern semantic technologies offer the means to make these heritage documents accessible by transforming them into a semantic knowledge base. Using techniques from natural language processing and Semantic Computing, we automatically populate an ontology. Additionally, all content is made accessible in a user-friendly Wiki interface, combining original text with NLP-derived metadata and adding annotation capabilities for collaborative use. All these functions are combined into a single, cohesive system architecture that addresses the different requirements from end users, software engineering aspects, and knowledge discovery paradigms. The ideas were implemented and tested with a volume from the historic Encyclopedia of Architecture and a number of different user groups.
OrganismTagger: detection, normalization and grounding of organism entities in biomedical documents
Abstract
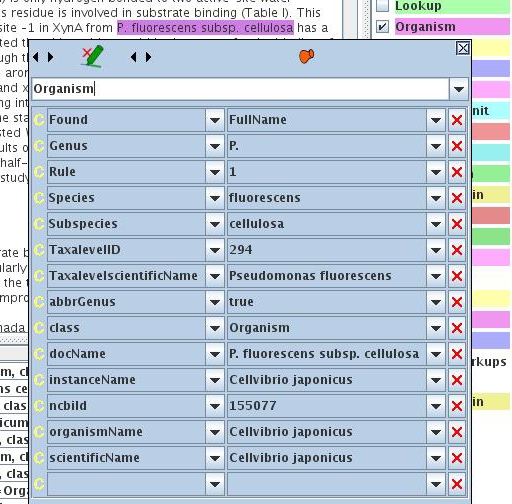 OrganismTagger example result annotation
OrganismTagger example result annotation
Motivation: Semantic tagging of organism mentions in full-text articles is an important part of literature mining and semantic enrichment solutions. Tagged organism mentions also play a pivotal role in disambiguating other entities in a text, such as proteins. A high-precision organism tagging system must be able to detect the numerous forms of organism mentions, including common names as well as the traditional taxonomic groups: genus, species and strains. In addition, such a system must resolve abbreviations and acronyms, assign the scientific name and if possible link the detected mention to the NCBI Taxonomy database for further semantic queries and literature navigation.
Results: We present the OrganismTagger, a hybrid rule-based/machine learning system to extract organism mentions from the literature. It includes tools for automatically generating lexical and ontological resources from a copy of the NCBI Taxonomy database, thereby facilitating system updates by end users. Its novel ontology-based resources can also be reused in other semantic mining and linked data tasks. Each detected organism mention is normalized to a canonical name through the resolution of acronyms and abbreviations and subsequently grounded with an NCBI Taxonomy database ID. In particular, our system combines a novel machine-learning approach with rule-based and lexical methods for detecting strain mentions in documents. On our manually annotated OT corpus, the OrganismTagger achieves a precision of 95%, a recall of 94% and a grounding accuracy of 97.5%. On the manually annotated corpus of Linnaeus-100, the results show a precision of 99%, recall of 97% and grounding accuracy of 97.4%.
Availability: The OrganismTagger, including supporting tools, resources, training data and manual annotations, as well as end user and developer documentation, is freely available under an open-source license at http://www.semanticsoftware.info/organism-tagger.
Intelligent Software Development Environments: Integrating Natural Language Processing with the Eclipse Platform
Abstract
 Software engineers need to be able to create, modify, and analyze knowledge stored in software artifacts. A significant amount of these artifacts contain natural language, like version control commit messages, source code comments, or bug reports. Integrated software development environments (IDEs) are widely used, but they are only concerned with structured software artifacts – they do not offer support for analyzing unstructured natural language and relating this knowledge with the source code. We present an integration of natural language processing capabilities into the Eclipse framework, a widely used software IDE. It allows to execute NLP analysis pipelines through the Semantic Assistants framework, a service-oriented architecture for brokering NLP services based on GATE. We demonstrate a number of semantic analysis services helpful in software engineering tasks, and evaluate one task in detail, the quality analysis of source code comments.
Software engineers need to be able to create, modify, and analyze knowledge stored in software artifacts. A significant amount of these artifacts contain natural language, like version control commit messages, source code comments, or bug reports. Integrated software development environments (IDEs) are widely used, but they are only concerned with structured software artifacts – they do not offer support for analyzing unstructured natural language and relating this knowledge with the source code. We present an integration of natural language processing capabilities into the Eclipse framework, a widely used software IDE. It allows to execute NLP analysis pipelines through the Semantic Assistants framework, a service-oriented architecture for brokering NLP services based on GATE. We demonstrate a number of semantic analysis services helpful in software engineering tasks, and evaluate one task in detail, the quality analysis of source code comments.
Believe It or Not: Solving the TAC 2009 Textual Entailment Tasks through an Artificial Believer System
Abstract
The Text Analysis Conference (TAC) 2009 competition featured a new textual entailment search task, which extends the 2008 textual entailment task. The goal is to find information in a set of documents that are entailed from a given statement. Rather than designing a system specifically for this task, we investigated the adaptation of an existing artificial believer system to solve this task. The results show that this is indeed possible, and furthermore allows to recast the existing, divergent tasks of textual entailment and automatic summarization under a common umbrella.
